 Author: Alex Pico aut, cre, Tanja Muetze aut, Paul Shannon aut, Ruth Isserlin ctb. Anything you can do using the graphical user interface of Cytoscape, you can now do with a single RC圓 function. Bioconductor version: Release (3.17) Vizualize, analyze and explore networks using Cytoscape via R. To view documentation for the version of this package installedĠ5. Functions to Access and Control Cytoscape. If (!require("BiocManager", quietly = TRUE))įor older versions of R, please refer to the appropriate The function can also work with just one dataframe.To install this package, start R (version Additional columns are imported as node and edge attributes into Cytoscape. 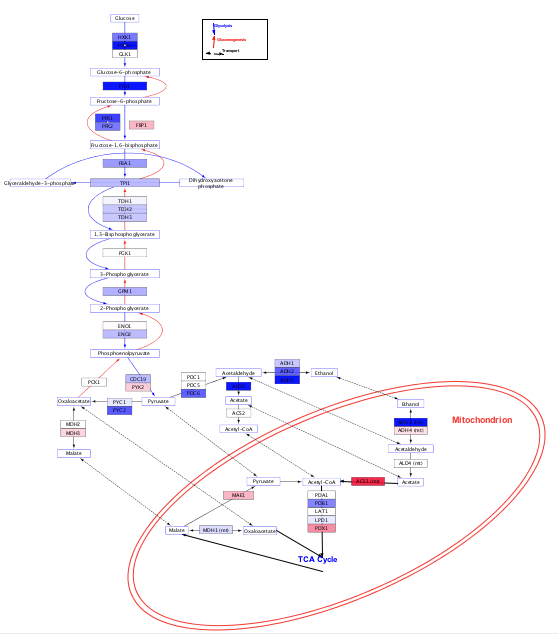
Nodes dataframe must include a column named Likewise, igraph and graphNEL objects can be created from networks (Ĭreate * FromNetwork), and dataframes from node and edge tables in Cytoscape (ĬreateNetworkFromDataFrames, two dataframes are accepted as arguments, one for nodes and one for edges. RC圓 can create networks in Cytoscape from either igraph, graphNEL or dataframe objects (ĬreateNetworkFrom *). Networks are a popular visualization option in R often implemented as graph models by a script author is presented with a series of intuitively named functions with obvious arguments. That is what will build the network, which usually one can make from gene lists putting them either in stringdb or with cytoscape with GO plugins and then build the relationships. With autocomplete in tools like RStudio, after just typing I am just thinking of how to put the relations of GO terms to get the edges and connection for making the graph. To simplify usage for common situations, we therefore also implemented specific functions for over a dozen of the most commonly used mappings (e.g., "NODE_FILL_COLOR", and mapping data structures. So it looks like you can perform all of your analysis just by using it. However, these functions are not simple to use, requiring knowledge of specific property names, like Blast2GO already supports InterPro, enzyme codes, KEGG pathways, GO direct acyclic graphs (DAGs), and GOSlim. (2003) 12 Cytoscape It is used for visualization and analysis of. In the current Blast2GO version this is the core type of functional annotation. (2012) html/edgeR.html 8 Blast2GO It is a tool used for functional annotation of. 
With these generic functions one can perform any of the hundreds of visual style mappings supported by Cytoscape, including new ones added in the future. Gene Ontology Annotation Content of this page: Blast2GO Annotation Rule This is the process of selecting GO terms from the GO pool obtained by the Mapping step and assigning them to the query sequences. Style.name and mapping arguments and sends them out via Blast2GO can be seen as a working platform to generate sequence functional data, and in fact many of the annotation aspects studied in the experimental part of this work could not have been addressed with any of the other tools. 
Two-way conversion with networks from \textit / mappings that takes a JSON data structure defining the mapping. Over 40 Cytoscape apps have implemented automation support so far, making hundreds of additional operations accessible via RC圓. Over 100 new functions have been added, including dozens of helper functions specifically for intuitive data overlay operations. RC圓 has been redesigned to streamline its usage and future development as part of a broader Cytoscape Automation effort. RC圓 is an R package in Bioconductor that communicates with Cytoscape via its REST API, providing access to the full feature set of Cytoscape from within the R programming environment. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
